新一代测序和桑格测序的差别
从原理上说,桑格测序与新一代测序(NGS)技术背后的概念是相似的。在NGS和桑格测序(也称为双脱氧测序或毛细管电泳测序)的过程中,DNA聚合酶会将有荧光的核苷酸逐个添加到正在延长的DNA模板链上。通过荧光标记识别每个掺入的核苷酸。
桑格测序与NGS的主要差异在于测序量。桑格测序一次只能对一条DNA片段进行测序,而NGS可以进行大规模平行测序,每次运行可同时对数百万个片段进行测序。这意味着这样的高通量过程一次可对数百到数千个基因进行测序。NGS还具有更强的探索能力,可以通过深度测序检测新型或罕见变异。

NGS相较于桑格测序的优势
NGS的优势包括:
- 更高的灵敏度,可检测低频变异1,2
- 周转时间更快,可检测的样本量更高3
- 全面覆盖整个基因组
- 检测限更低4,5
- 可进行样本多重分析,通量更高
- 能够同时测序数百到数千个基因或基因区域
如何选择NGS与桑格测序
了解每种方法的优点与局限性,选择最适合您的方法。
“利用桑格测序,我们只能看到有限的DNA快照……NGS及其大规模平行测序使我们能够分析每个样本的数万乃至数十万条read。”
Michael Bunce博士
澳大利亚科廷大学教授,TrEnD实验室主任
桑格测序与NGS的比较
| 桑格测序 | 靶向NGS | |
|---|---|---|
| 优势 |
|
|
| 挑战 |
|
|
* 探索能力为鉴定新型变异的能力。
† 突变分辨率是指鉴定的突变大小。NGS可以进行大的染色体重排到单核苷酸变异的鉴定。
‡10 ng DNA用于桑格测序将产生~ 1 kb,用于靶向重测序将产生~ 300 kb(利用TruSeq Custom Amplicon流程,250 bp的扩增子长度 × 1536个扩增子)
何时使用NGS?何时使用桑格测序?
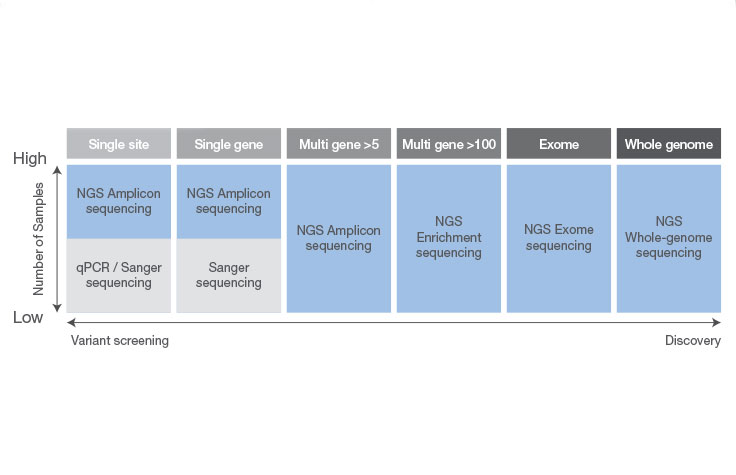
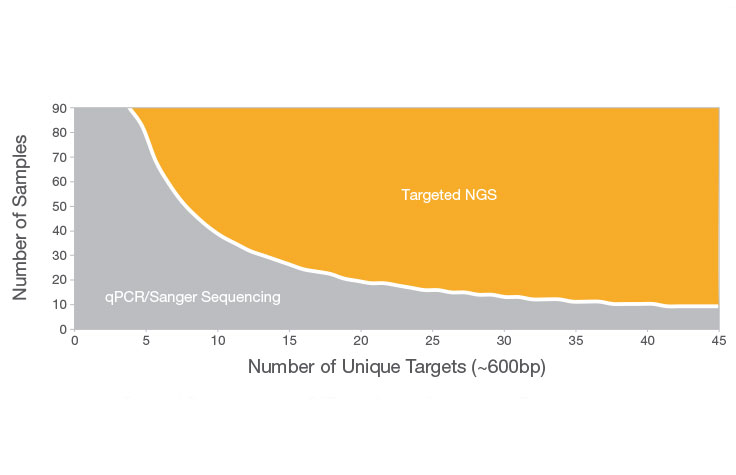
对少量样本或少量基因组靶点(≤20个)的小DNA区域进行测序时,桑格测序是一个很好的选择。其他情况下则靶向NGS更能满足您的需求。NGS可让您经济有效地筛查更多的样本,检测基因组目标区域的多个变异——如果使用桑格测序开展这些分析则昂贵且耗时。
观看动画,了解简单易用的Illumina NGS技术如何补充您的桑格测序工作。
从桑格测序转换到NGS
iSeq 100测序系统能让您的实验室比以往更简单、更经济的使用NGS的强大功能。以下示例说明了该仪器如何适应常用的NGS工作流程。
相关解决方案
靶向重测序

通过靶向重测序,可以分离出一部分基因或基因组区域并进行测序,从而节约实验室资源。深入了解靶向重测序。
全基因组测序

全基因组测序可以提供遗传变异的全面视图,非常适合探索应用。深入了解全基因组测序。
NGS vs. 桑格测序
更多资源
参考文献
- Jamuar SS, Lam AT, Kircher M, et al.Somatic mutations in cerebral cortical malformations.N Engl J Med.2014;371(8):733-743.
- Rivas MA, Beaudoin M, Gardet A, et al.Deep resequencing of GWAS loci identifies independent low-frequency variants associated with inflammatory bowel disease.Nat Genet.2011;43(11):1066-1073.
- König K, Peifer M, Fassunke J, et al.Implementation of amplicon parallel sequencing leads to improvement of diagnosis and therapy of lung cancer patients.J Thorac Oncol.2015;10(7):1049-1057.
- Shendure J and Ji H. Next-generation DNA sequencing.Nat Biotechnol.2008;26(10):1135-1145.
- Schuster SC.Next-generation sequencing transforms today’s biology.Nat Methods.2008;5(1):16-18.